My work focuses broadly on understanding how biochemical pathways evolve. Metabolic pathways underlie various cellular processes, yet it is unknown how pathways are constructed and by what genetic, biochemical, and biophysical mechanisms pathways evolve.
What happens to genomes and biochemical networks when species hybridize?
We are interested in understanding what the genomic and metabolic consequences of hybridization between Asclepias syriaca (the common milkweed) and Asclepias speciosa (the showy milkweed). These species form a wide hybrid zone in central North America and we are studying the consequences of this hybridization through common gardens, greenhouse, and lab experiments. Find out how to contribute to this project with the milkweedflower.org
How is functional evolution constrained?
A critical question in molecular evolution is how do proteins gain novel function. However, even after the molecular mechanisms for functional evolution are discovered, many questions still remain. Were other mechanisms for the same functional shift possible? How was the functional outcome influenced by the starting protein sequence? Fortunately, convergent evolution of proteins offers parallel natural experiments of functional evolution. With phylogenetic and biochemical techniques, I am studying a convergence in corticosteroid synthesis in the rodents and primates. The genetic mechanisms in the convergence events are different and I am currently studying how the biochemical mechanisms in the two groups compare.
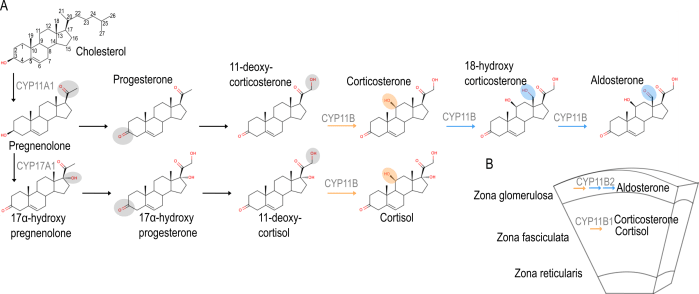
Corticosteroid synthesis pathway in humans
How does novel function arise in metabolic pathways?
Understanding how novel function evolves requires information about each step in the link between genotype and phenotype, from the function-changing mutations, to its effect on phenotype and fitness. Furthermore, understanding gain-of-function mutations is even more challenging as they are rare and easily lost in natural populations. However, these mutations are critical for evolutionary innovation. I use a variety of genomics and biochemical techniques to try to understand how a novel glucosinolate was produced in the Arabidopsis relative, Boechera stricta (Prasad, Song, Olson-Manning et al. Science 2012).
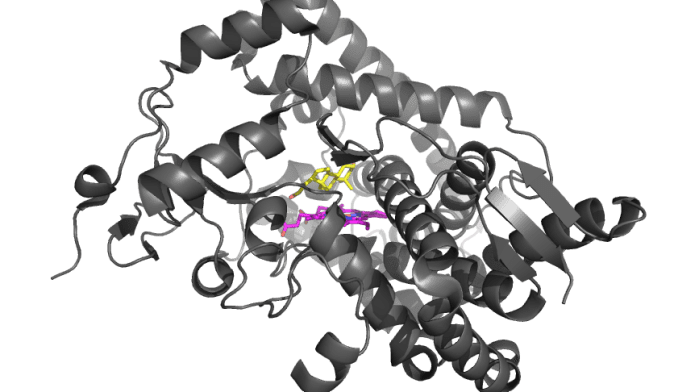
How do network properties of an enzyme influence how selection acts?
An important finding in the post-genomics era is that genes evolve at very different rates. One leading theory is that the rate of evolutionary change in enzymes may be driven largely by their network context. One such network characteristic is the amount of control an enzyme has over the amount of product produced, or flux control. With genetic experiments I have found that enzymes with majority flux control are more likely to be targets of natural selection (Olson-Manning et al. 2013 MBE, Olson-Manning et al. 2015 G3: Genes|Genomes|Genetics) that evolution may predictably act on certain parts of pathways based on their network properties.
Mate preference in butterflies?
In collaboration with Dr. Erica Westerman at the University of Arkansas, Dr. Marcus Kronforst at the University of Chicago, and the Butterfly House and Marine Cove we are conducting a series of behavioral assays of mimetic butterflies. For more information, see Dr. Westerman’s research page.